Question and answer (Q&A) as well as frequently asked questions in Magnetic Resonance.
Q&A for Principles of Magnetic Resonance (Spring 2020)
Question: I am dealing with travel, social isolation, etc… related to the coronavirus. How is this course dealing with this global pandemic?
First, don’t worry AT ALL about this course and deal with your immediate personal health and family health issues. You can ALWAYS deal with the requirements for this course at a later date and any students that do not have the resources or bandwidth or are dealing with issues related to the global pandemic will receive an incomplete and can work with Prof. Yarger to finish the requirements for this course in a future semester (at a later date). Also, we are trying to move as much of current and future online coursework for PMR (and all online course that Prof. Yarger teaches) to asynchronous ‘courses’ and material.
Asynchronous learning is a student-centered teaching method that uses online learning resources to facilitate information sharing outside the constraints of time and place among a network of people. In my opinion, this is a method we need to implement immediately for Principles of Magnetic Resonance, Spring 2020. (In many ways, we already have implemented this ‘self-pace’ method)
I will be updating my blog and PMR website at biopchem.education with more details (by the end of this week). I have received several emails from students over the past several days and instead of addressing them individually, I am providing some key information below:
- PMR will be completely online and primarily asynchronous for the remainder of the semester.
- The only components that will not be asynchronous are the peer reviews for PMR Unknown #2 and Project #2. All due dates and instructions are provided in the assignment tab on Canvas. However, outside of the peer reviews, which are optional, the final two evaluation components for this course (unknown #2 and project #2) and be turned into canvas anytime for evaluation and all resources for learning principles of magnetic resonance made available at biopchem.education for students to use whenever is convenient for them.
- Zoom will be used for office hrs (Wed. 3:15-4:15 pm) and students can email to setup a zoom appointment outside of this standard time. This will NOT be a lecture and is NOT required to attend. It is an optional time for students to communicate online with Prof. Yarger. Asynchronous communication with Prof. Yarger and peer-to-peer is being setup now and will hopefully take the place of this scheduled weekly office hr. More information on this shortly.
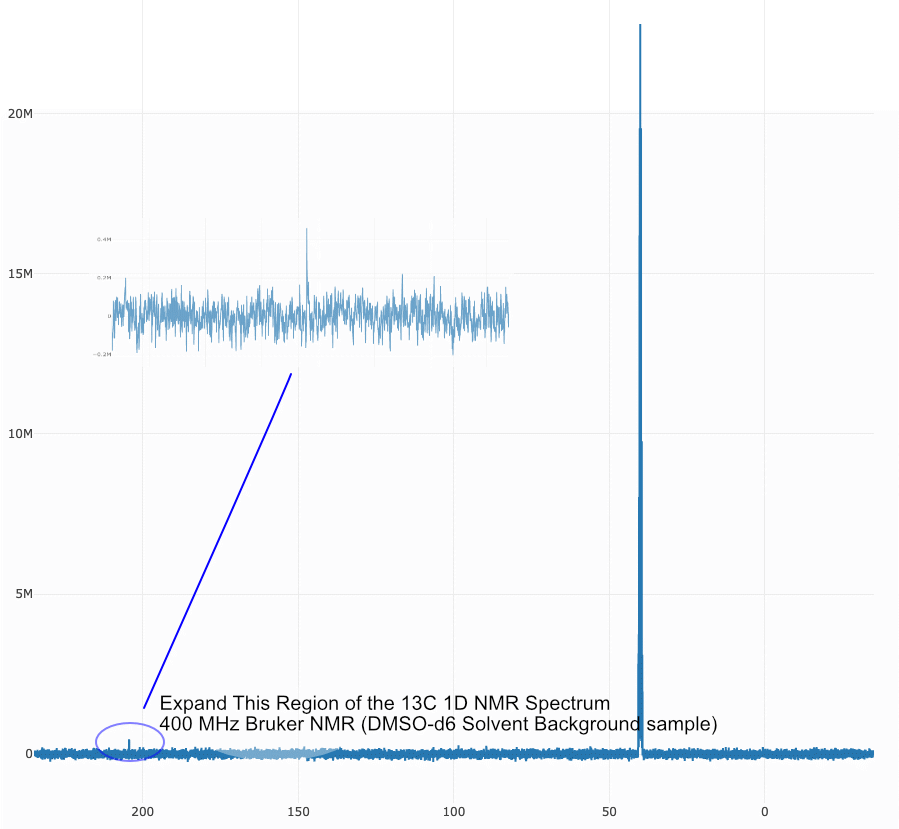
Question: In unknown samples there is a peak in the 13C 1D spectrum at ~210 ppm. This peak doesn’t show up in any of the hetcor 2D experiments (e.g. 1H-13C HSQC or HMBC). Why?
Answer: Looking at a bunch of the solvent samples and standards, you can often see this same peak. Hence, it appears to be an artifact and not a resonance coming from your sample. This seems to appear only on the 400 MHz Bruker system and the NMR operations manager has been notified to better diagnose the instrument issue. However, this is a great lesson in ALWAYS running a background solvent sample (with everything EXCEPT your sample in the same type of NMR tube, and also running a standard, calibration or know sample) to insure the instrument is working properly.
Question: For my assigned unknown organic compound, there is a set of data already collected on spintropy. What NMR data was collected and is this all the data I need to solve the molecular structure of my unknown organic compound (between the empirical formula CHO)?
Answer: I ran a standard ‘MRRC Structure’ set, which consists of a 1H, 13C, COSY, HSQC and HMBC NMR data. This is the typical set of NMR experiments run to determine molecular structure for standard organic molecules (under 1000 g/mol molecular weights). However, it is common that ‘other’ NMR experiments are useful or even needed to determine molecular structure definitively, especially if there are other heteronuclei besides Carbon and Oxygen (e.g. Nitrogen, halogens, phosphorus, etc). Also, there are many other NMR experiments that help elucidate molecular dynamics and chiral molecular confirmations.
Students can request additional NMR experiments for their unknown sample. All they need to do is email Prof. Yarger at yarger@biopchem.education and explain what additional NMR experiment(s) they want to have acquired and WHY. The justification of why they want additional NMR experiments is critical to the request being accepted or denied.
As an example, one student asked for a DEPT135 experiment to be run on their unknown. Their justification is that this is needed to help with carbon assignment of their unknown. The request was denied because the student already had a phase sensitive HSQC was run with multiplicity editing on their sample which provides the information they were requesting from the additional DEPT135 experiment. The student was pointed toward the following information on this topic:
Magritek Blog on Multiplicity Edited HSQC.
As another example, one student asked for a NOESY experiment to be run on their unknown. They justified this as an experiment that would help them with molecular confirmation because of molecular long range through space contacts that NOE’s provide. Their molecule has a chiral center and the student is trying to determine information about absolute molecular confirmation. This request was approved and the 2D NOESY was run on the 500 MHz Bruker NMR overnight. Data was automatically uploaded to Spintropy. A blog related to this topic is provided below (pdf format).
Question: I am having an issue with the ADEQUATE NMR data, can you help? I think the offset is incorrect.
Answer: The ADEQUATE takes awhile to wrap your head around!! and I didn’t realize that the auto processing didn’t get the offset/ref correct, BUT it doesn’t surprise me!! The referencing gets messed up in auto-processing 10-20% of the time, especially in cases of low SNR.
Frequently Asked Questions (and answers)
Question: Don’t I need the exact molecular weight and/or Mass Spec data as well as functional group data from IR to determine the molecular structure of an assigned organic unknown compound?
Answer: The short answer is YES. To unequivocally determine molecular structure it really helps and is often required that you have Mass Spec data, IR data and elemental analysis data. However, NMR is the PRIMARY technique that really unequivocally proves molecular bonding connections and for A LOT of cases NMR can often be used exclusively to determine molecular structure or at least GREATLY reduce the number of potential molecular configurations that are consistent with the NMR data/interpretation. Because this course is focus exclusively on magnetic resonance, you will only be given NMR data and we feel that it is really instructive to see how far you can get toward molecular structure elucidation with ONLY magnetic resonance experiments.
Test Comment… Students, please use this Q&A page to ask questions and provide comments.
LikeLike