Determining the molecular structure (and/or the molecular dynamics) of chemical compounds (typically organic compounds or organo-metallic compounds) is the most common use of Nuclear Magnetic Resonance (NMR) spectroscopy. In ‘Principles of Magnetic Resonance’ (PMR), we first like to practically use NMR and then use this practical experience to build in a fundamental understanding (principles). Hence, it is our advise that one of the first things a person should do is get some ‘hands-on’ NMR experience and there’s not a more common place to start than structure elucidate of organic molecules dissolved in a (deuterated) solvent. The assignment for students is to determine the molecular structure of an unknown organic compound dissolved in a specific (common) solvent (typically deuterated chloroform, CDCl3 or deuterated water, D2O). All data can be found on the course spintropy account (ASU_PMR_Class).
PMR Unknown Cmpd #1 (Jan/Feb 2020)
Details about the NMR solvent, tube, sample, etc is given as comments on the Bruker topspin data directly. Most of the ‘unknown’ samples were dissolved in CDCl3 w/ 1% TMS (v/v) and all samples were run in 5 mm Norell Sure Seal NMR tubes on the 400 MHz and/or 500 MHz Bruker Avance NMR system(s) in the MRRC at ASU (ISTB1 L2-63). All ‘PMR Unknowns’ assigned to students for molecular structure elucidation (Jan-Feb, 2020, PMR # 19, 20, 22, 23, 24, 25, 27, 29, 32, 34 and 36) are between 10-15 Carbons, 10-25 Hydrogens and 1-5 Oxygens (C10-15H10-25O1-5). All molecular compounds only contain hydrogen, carbon and oxygen (HCO compounds). It is really hard to EXACTLY solve the molecular structure of an unknown organic molecule without some additional data. For example, having as least 1 of the following is often required or at least very very helpful to solve the molecular structure:
Mass Spectroscopy (MS), Infared (IR), Melting Pt, Elemental Analysis.
However, as this class is focused on magnetic resonance, it is often very instructive to try and solve the molecular structure using only NMR experiments to predict and verify a molecular structure. In most cases, for such low molecular wt organic compounds this is reasonable. And even if an exact structure can’t be proven, it is always possible to greatly limit the potential molecular structures that are consistent with basic and advanced NMR experiments.
After the initial ‘first draft’ is due, additional information and discussion about the PMR Unknown Organic Compounds will be provided at the following link:
Additional PMR Unknown Cmpd #1 Information
It is becoming common for complete automation of data collection by NMR spectrometers. Both the ASU MRRC Bruker 400 MHz Avance III NMR and the Bruker 500 Mhz Avance Neo NMR system have sampleXpress autosample systems and allow users to pick from a common set of NMR experiments using ‘icon NMR’ (Bruker automation software). All a user has to provide is the deuterated solvent, sample name and what common NMR experiment or set of experiments they want collected. One of the sets is called ‘MRRC_Structure’ and consists of running 1H, 13C, COSY, HSQC and HMBC NMR experiments on the sample. Besides have this data available on Spintropy, a zipped version of this Bruker NMR data is provided below (so that anyone can download the data for educational purposes). Once students start to analyze their data, they can request ADDITIONAL NMR experiments be performed on their unknown sample. However, a justification must be provided and the instructor has to approve the request as reasonable and justified.
PMR Unknown Cmpd #2 (March/April 2020)
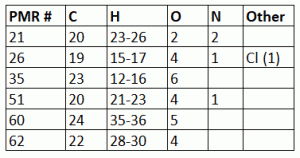
Details about the NMR solvent, tube, sample, etc is given as comments on the Bruker topspin data directly. Most of the ‘unknown’ samples were dissolved in CDCl3 w/ 1% TMS (v/v) and all samples were run in 5 mm Norell Sure Seal NMR tubes on the 400 MHz and/or 500 MHz Bruker Avance NMR system(s) in the MRRC at ASU (ISTB1 L2-63). Data for students in PMR can be downloaded from the ASU_PMR_CLASS spintropy account. Unlike Unknown #1, this time students are NOT assigned specific unknowns. Instead students can pick from any of the PMR Unknown #’s 21, 26, 35, 51, 60 or 62. All of these unknown organic molecules are pharmaceutical (drugs) compounds used in primarily for treating health issues. A table of C, H, O, N and any heteronuclei are provided (PMR Unknown Cmpd #2 List). The number of hydrogens (H) is given as a range which include all possible ionic and hydrogen exchangeable states of the unknown molecule. A standard basic set of 1D and 2D NMR experiments have been done on all these cmpds, including 1H, 13C, 1H-1H COSY, 1H-13C HSQC and 1H-13C HMBC NMR. Also, most of these samples will also include a 1H-13C HSQC-TOCSY and 1H-1H NOESY. Also, any of the cmpds that contain nitrogen will have a 1H-15N HSQC. For a limited number of these compounds we are working to get 1H-13C ADEQUATE experiments (instrument time and sample concentration limit the ability to collect this carbon correlation experiment on all samples).
Links to some websites, videos, books, journal articles and general information that is useful when learning how to solve molecular structure from NMR and related experimental data:
- Structure Determination Using Spectroscopic Methods – Reich (U. Wisc.)
- U. Ottawa NMR Blog – Glenn Facey.
- Chem Info Website
- Web Spectra (UCLA)
- Organic Structure Elucidation @ Notre Dame
- Emery Pharma -Step-by-Step Guide to 1D & 2D NMR Interpretation
Links to videos, tutorials and information on data processing and analysis using Bruker TopSpin software:
Are you still running PMR samples as of Mar 23 or are you only running those requested? I ask because I don’t see data for PMRs 26, 35, and 51. I do see data for PMR 61 however which isn’t in the cmpd #2 list.
LikeLike
All preliminary data for Unknowns #2 have been run. In fact, many of the basic 1H, 13C, COSY, HSQC, and HMBC experiments were run back in January. Hence, you may need to search for the data in Spintropy, because only the past 30 days (or less) of data is shown by default. However, you have access to ALL the past data collected for ASU_PMR_Class on spintropy (over the past year). Because of the automated nature of the NMR instruments at ASU, I should be able to run requests for students, as long as they don’t require major time spent in the lab. Please email me at yarger@biopchem.education if you have any specific requests. As for PMR61, the initial MRRC Structure data has been completed on this sample quite some time ago, just do a search under spintropy and you should find it. If not, let me know and I can help you search and find it.
LikeLike
Awesome, thank!
LikeLike
What’s the difference between a 1, 1 – ADEQUATE and a 1H-13C H2BC? Both descriptions I read give me the same answer but I see there are more peaks in my adeq11 for pmr21 and I don’t know why.
LikeLike
The ADEQUATE is similar in information to an INADEQUATE and primarily gives Carbon-Carbon correlation information. However, it does it through protons for sensitivity reasons. A good online reference is: https://www.bruker.com/fileadmin/user_upload/8-PDF-Docs/MagneticResonance/Events_NMR/SMASH2016/1_1-ADEQUATE_Exclusively_Affords_1JCC_Correlations.PDF
LikeLike
Can I get an HMBC on the new, more concentrated PMR21? The old HMBC is contradictory with the new 1, 1 – ADEQUATE.
LikeLike
Unfortunately, the automated NMR’s are not in service at the moment and policies are changing quickly because of COVID-19. We can try to collect more data in the next few weeks, but I’m guessing this is not soon enough for what you need. The PMR21 sample has been made several times in several concentrations and even with 2 or more solvents. The higher concentrations are required for insensitive experiments like an ADEQUATE experiment. However, high concentration can lead to aggregation and large changes in chemical shift and other issues that general is better to avoid by running more dilute solutions of the molecule of interest.
I understand that there are some differences between NMR spectra of several of the PMR21 samples. This can be caused by several things including, (i) high concentration interactions (dimerization, oligomers), (ii) different solute-solvent interactions, (iii) degradation over time, etc. I understand this can make things more difficult to interpret and often more data and showing reproducibility is useful to insure (iii) isn’t playing a significant role.
LikeLike